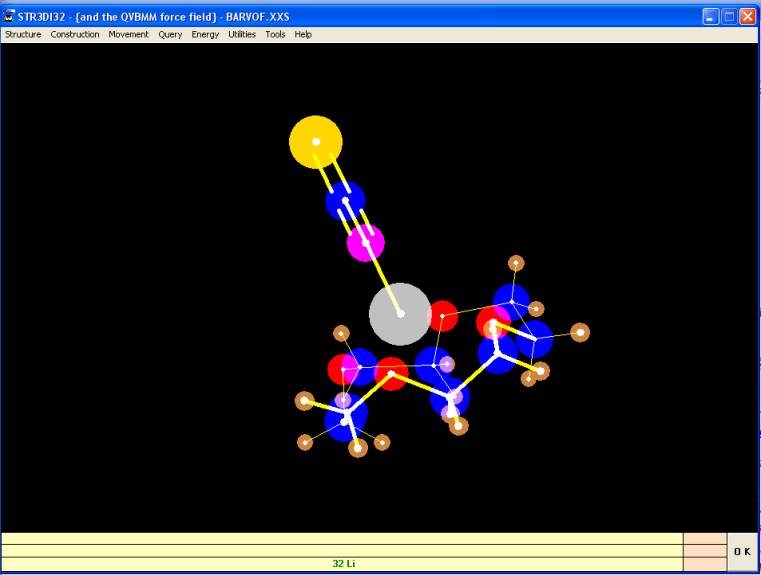
BARVOF, ionic bonds not shown
The full-featured (no nags, no limits in capabilities) demo version is available for FREE! However, the program is still copyrighted to Vernon G. S. Box
StruMM3D version 9.0.0.38 is now available and has been greatly improved. The molecular recognition algorithms have been updated, the QVBMM Force Field has been tweaked to make it faster, and several new features have been introduced.
By now you would know about the Molecular Log that StruMM3D uses to create an inventory of the molecules accessed, and this log file can be searched from withing StruMM3D to find whatever you need. In version 8.0.0.14 we also introduced a more "visual" way to search for target molecules. If you copy the screen/displayed molecule to the clipboard a copy of the JPG or BMP file is also automatically stored in the working directory. You can use that graphics file in your publications or documents. The collection of BMP/JPG/GIF files can become a visual record of your files, via their thumbnails.
If you have kept the graphics file in the same directory as the coordinate data file, then when you right click on the graphics file and select the "Open With" option to use StruMM3D to open the file, the molecular modeling exercise will be launched. So now you have a visual way of identifying molecules and loading them into StruMM3D, as well as the older Molecule Log method.
A historical/developmental look at the rise of StruMM3D follows.
StruMM3D version 7.2.0.37 became the first, ever, molecular mechanics program that enabled the user to examine every aspect of a transition state structure. StruMM3D automatically identifies transition state bonds, even in x-ray crystallographically determined structures. StruMM3D has never used human-generated connectivity tables. This is a boon to molecular voyeurs and crystallographers alike. All chemists will be delighted by these new features in StruMM3D, especially since the transition state models can be systematically modified to reveal energy trends.
From the release of version 7.2.0.45, users will enjoy the full capabilities of our prized data file conversion utility for free. So go ahead, just launch StruMM3D, load any file of coordinate data, from most molecular modeling suites (CSD, PDF, CIF, XYZ, MOL, etc.), and you'll have a structure to examine. Unless, of course, the structure data file has flaws, which is common because they are so complex, unlike StruMM3D's files.
StruMM3D version 7.2.0.51 marked the successful attainment of one major step in StruMM3D's quest to finding global molecular energy minima, and new code for calculating bond orders. Version 7.2.0.54 featured a re-worked algorithm for the automatic detection of functional groups, better program stability, and more conservative memory/energy usage.
The release of StruMM3D version 7.3.0.1 marked an important milestone. This version sported a new colour scheme for atoms, which carries more structural stereo-electronic information. Re-designed working windows, more crash-proofing protection (from bad data and ill-advised user input), and faster graphics rounded out the list of obvious improvements. There are others too, but we can't elaborate here.
Many theoreticians boast that their computer modeling method/tool will generate good simulations of organic molecules, but their methods cannot, and do not, inform the user of some of the basic stereo-electronic features of these molecules. So these simulations are just not scientifically useful to someone searching for information. Imagine looking at the x-ray crystal structure of a transition metal complex in which the metal has 7 ligands. Can your molecular modeler help you to determine what the oxidation state of the metal is? Is there any way to even ask this questions using this molecular modeler?
In StruMM3D version 7.3.0.3 the advanced structure recognition algorithms used will enable you to examine any transition metal complex, with any coordination number, to identify the oxidation state of the metal, and hence to identify one electron single bonds in that complex. Since transition metal based catalysts are now very important reagents in organic chemistry, your molecular modeler of choice ought to be StruMM3D, if you really wish to understand how these catalysts work. StruMM3D not only generates excellent molecular simulations, but it also reliably and transparently informs the user about all of the stereo-electronic features in the molecular model.
StruMM3D version 7.3.0.33 boasts some newly developed structure recognition and manipulation algorithms that have never before been implemented in a molecular modeling program. In particular, these developments will greatly facilitate the simulation of organometallic compounds. This new sophistication will enable users to easily perform molecular modeling studies not easily done in the past.
StruMM3D version 7.3.0.53 now allows you to perform a lightening fast search of all of the CIF, PDB and ENT files that are stored in the \Str3Di folder, on your computer, so that you can identify molecules you are looking for. This feature supplements the Molecule Log feature that allows you to search for molecules by functional groups, or by molecular composition.
StruMM3D version 7.3.0.60 and all newer versions will be compiled in the Windows 10 Pro 64 bit environment, with all the compiler switches set to produce the fastest, and leanest, version.
====================
StruMM3Dis poweredby the mostsophisticated molecular mechanics force field ever developed, The QVBMM force field. StruMM3D's advanced structure recognition and analysis algorithms enable the user to input molecular structure data by using only atom types and atomic coordinates (ensuringthat faulty user-generated atom connectivity list never introduce structural errors into the molecular modeling exercise) and enable the user to examine any, and all, stereo-electronic effects in the molecule.
====================
StruMM3D generates the best molecular simulations of single molecules and docked molecular clusters, of any molecular mechanics program currently available. If you want to do serious, sophisticated molecular modeling, like working with biomolecular clusters, including chelated metal atoms, and the many other things that the student versions of software packages cannot do, then you will want to use StruMM3D.
If you want to do modern molecular modeling, in which molecular structural features are generated in direct response to the structural and stereo-electronic features of the molecule being simulated, and NOT by inflexible templates, then consider using StruMM3D.
Try using one of the other molecular modelers to search a large complex protein structure for important features like non-delocalized amide linkages! Only StruMM3D can do that successfully.
The advanced structure analytical algorithms of StruMM3D have been exhaustively tested, and shown to be vastly superior to others, in detecting subtle stereo-electronic effects embedded in diffraction generated coordinate data, like delocalization, aromaticity and anomeric effects. No other molecular mechanics based program can help you to understand the interplay of structure and reactivity because they do NOT perform their calculations with explicit inclusion of lone pair interactions. For example, the ONLY molecular mechanics program that can predict the relative nucleophilicities of the hydroxyl groups in a polyol is StruMM3D.
Did you know that the vast majority of quantum mechanical theoretical studies start with molecular models generated by molecular mechanics?
And that even when a low-end MM program is used to make the initial model, the QM manipulations hardly ever result in any change in the structure?
Did you know that it is almost impossible to generate a realistic molecular model, ab initio, using any QM method?
And, when you do your simulation, did you use the right/correct basis-set, the best set of approximations, to get the best model?
====================
And, while we are here, PLEASE remember that........
StruMM3D is a 32 bit program. Tell us about any compatibility issues that you might encounter because we can help you to get going.
We have heard about User Account Control (UAC) issues that you might have had in the past, and these should no longer occur. StruMM3D is totally compatible with Windows Vista, 7, 8 and 10. You should not need to execute StruMM3D by using the “run as an administrator” option. However, if StruMM3D does not start the first time because of the way your computer is set up, just right click on the file StruMM3D.Exe in the directory/folder \Str3Di, and select “Run as an Administrator”.
Most emulators that run Windows in Apple Macs will run StruMM3D almost as fast as it runs in a PC with Windows.
We have added a short tutorial on drawing molecules using StruMM3D at the StruMM3D FAQ site. Take the time to find out more about StruMM3D and remember to visit the StruMM3D FAQfor hints, user/program tips and additional help.