Consultants in chemistry and chemical engineering, actively involved in computational, experimental, and environmental organic chemistry.
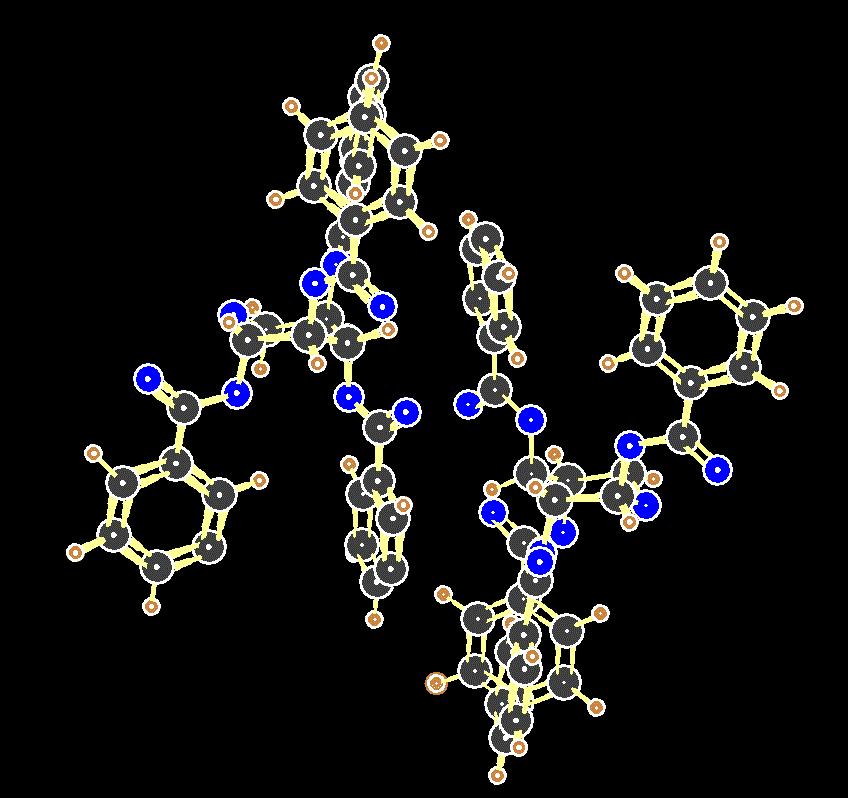
Two molecules of the tetrabenzoate of xylopyranose in the unit cell of its crystal. Note that the counter-intuitive axial conformation is favoured in the solid phase.
Find out about StruMM3D's molecular models, how you can generate them, and how you can use them in your daily tasks, presentations, reports and publications.
StruMM3D, The MOLECULAR MODELER
StruMM3D is now tightly integrated with its file importation utility, FileConv, enabling it to read many structure data file formats, and boasts new structure display features and options. Starting with version 4.6.0.1, StruMM3D had a totally revamped structure-energy minimization routine, to provide more accurate structure energy minima. New algorithms link your computer's performance to the structure energy minimization mode used.
StruMM3D was written and developed by a practicing synthetic organic chemist, who ensured that the simulations generated by StruMM3D were consistent with experimental organic chemistry. Many of the unique features of StruMM3D were developed because of the familiarity of the author with the subtleties of organic chemistry, and to ensure its compatibility with organic chemistry. For example, unlike most other molecular modelers, StruMM3D always generates a molecular simulation that visually presents the bond length and bond type data because this data is invaluable.
StruMM3D NEWS
· improved user friendliness
· improved molecular analytical and recognition features
· improved automated features, you NEVER have to write a connectivity list/table and so you never have to worry about making mistakes there.
· improved structure handling
· improved structure energy minimization
· improved system memory usage
· improved file name recognition (especially for long file names) for atomic coordinate data files, so enabling you to give these files usefully descriptive names.
· improved HTML-based molecule log that enables you to quickly retrieve structures you have saved, by way of the description of the molecules that you entered.
· StruMM3D enables you to view "slices" of large molecules. An excellent way to review the base pairs in a DNA double helical stack and other pi-stacked systems.
· StruMM3D enables you to dock two, or more, molecules. The structure energy of the cluster can be minimized, and all interaction energies can be measured. No other molecular mechanics based program can do this
· Internet based, as well as locally accessible, help files
· New sound prompts and flourishes
Rigorously tested under Windows 95/98/ME/NT3.5/NT4/2000/XP/Vista/7/8/8.1/10
StruMM3D can automatically recognize and model allenic systems, carbocations, and interesting systems like vinyl cations, protonated aldehydes and ketones. We have increased the number of atoms/lone pairs that will be handled by the QVBMM force field (in one logical molecular model) to 1000, so enabling the user to simulate really complex large molecules, like proteins and DNA/RNA oligomers.
We have developed the StruMM3D into an easy-to-use, very versatile, efficient and dramatically powerful software package. This software can deliver unique, insightful molecular modeling experiences to all organic chemists and crystallographers. Undergraduates, high school teachers, professors, and advanced researchers will enjoy using this software to enhance their understanding, and presentations, of organic molecules. The Str3Di molecular modeler will bring molecular modeling from the World Wide Web into your office, lab., or living room. Find out more about the Str3Di MOLECULAR MODELING PROGRAMS.
StruMM3D has been used to produce data for several scientific publications, in highly regarded refereed scientific journals, thus assuring you of its rigor and acceptability. Indeed, these papers will demonstrate, and validate, the our assertion that the Str3Di molecular modelers can provide solutions to molecular structure-reactivity problems that cannot be addressed by many other molecular modeling programs. Further, StruMM3D's uniquely powerful ability to examine, and extract information from, x-ray crystallographic coordinate data is also highlighted in these papers.
StruMM3D is powered by the new QVBMM MOLECULAR MECHANICS FORCE FIELD - recently described as ".the next generation of molecular mechanics force fields.." will do everything that any modern molecular mechanics, or semi-empirical molecular orbital, program can do. This force field is vastly superior to all other molecular mechanics force fields..
If you are unsure about the benefits of using an advanced molecular mechanics force field instead of quantum mechanical calculations then read this note on ab initio calculations to assure yourself that quantum mechanical calculations are not as "ab initio" as they are touted to be. These quantum mechanics calculations are just as approximate as molecular mechanics calculations, and are often incorrect. Did you know that ALL of these QM calculations start with molecular mechancis generated model? None of these QM programs can generate a viable structure from “scratch”.
For example, StruMM3D will -
dock molecules under the influence of their stereo-electronic interactions - structure energy minimization and modeling of dimers, trimers, and oligomers is a longstanding problem for other molecular mechanics programs
predict the selective nucleophilic reactivities of molecules that have several nucleophilic centers - a very valuable feature if you are working with sugars or other complex biomolecules
thoroughly analyze the structural features of molecules (experimentally determined by the diffraction methods, or generated by other theoretical methods). Many of these structures, which include important biomolecules, are riddled with unrealistic geometrical features that are not detected by the other currently available molecular modeling programs. You really will be surprised by the frequency with which unrealistic geometrical features are present in molecular models that are used by bio-organic chemists for important structure-activity studies. There are several websites that show "pretty pictures" of organic molecules, many of whose bond length and bond angle features are severely flawed.
generate structure energy minimized molecular models that have more plausible geometrical features than those generated by most other Molecular Mechanics programs - this is possible because of QVBMM uses ALL of the basic stereo-electronic effects that apply to all molecules, all the time.
rapidly search any molecular structure, even large complex molecules, for instances of unique, or common, structural features and functional groups - you won't understand how valuable this feature is until you try using any other software to search a crystallographic structure for an unusual entity, like a truly non-delocalized example of a pi-system that is "normally delocalized" (such as twisted, non-delocalized, amide units in a large complex structure).
automatically detect ground-state delocalized pi-systems - a boon to organic chemists, photochemists, and to researchers working with molecules that have extensive pi-systems. There are many molecules whose structures suggest that they should obey Huckel's Rules, but which, in fact, do not. The definitive way to confirm this is to examine the crystal structure data of that molecule, or a close relative, with StruMM3D.
Str3Di SUCCESSES
Small Data File and program size
We have had some whimsical comments on the very small size of the StruMM3D program (525 K), and its data files (most are less than 2 Kbytes and are also the smallest from any molecular modeling program). The program is small enough to be run from a floppy diskette! The size simply gives you a clue as to the sophistication of the algorithms used in the program. StruMM3D does not need the reams of data the old-styled multi-megabyte molecular modeling programs do (which they also read in from large data files). StruMM3D synthesizes much of the needed data from some basic scientific facts and new relationships, using some elegant algorithms.
Short Bonds? Weird Data?
One of the real hurdles that remain in modern molecular modeling is the continued use of structure coordinate data files in which bond types are explicitly specified. The creator of these data files can introduce many fatal errors into the structure. For example, a particular carbon - carbon bond in a molecular model can be designated a single bond, even though the length of that bond is less than that of a carbon - carbon triple bond! This type of flawed data is frequently encountered in coordinate data files that are generated by programs that convert 2-dimensional (2D) structures into 3D structures, and, unfortunately, these errors are found, occasionally, in X-ray crystallographic coordinate data files (put there by humans).
StruMM3D is the ONLY molecular modeling program that is sophisticated enough to automatically determine the types of all of the bonds in any molecular model, using only the coordinate data. StruMM3D does not use a connectivity list that is generated externally. StruMM3D cannot be induced to present a bond with an order that is not consistent with that embedded in the coordinate data. StruMM3D will automatically detect these weird bonds in structures that are flawed, and will immediately inform you of the presence of these "short bonds" in your molecular model, so enabling you to take appropriate corrective action.
In contrast, other molecular modeling programs rely on externally generated connectivity lists (made up by the user) and will not question that data. They will simply present to the user the information that is embedded in the (possibly flawed) data. Rubbish in, rubbish out! Of course, if your molecular modeler does not draw your attention to a serious structure flaw, you might believe that the structure is OK and continue to derive useless data from that flawed structure.
Visit our Exorga's DownLoadables site to find out how to acquire our molecular modeler for WINDOWS 95/98/ME/NT/2000/XP/Vista/7/8/8.1/10.